CPTR Newsletter – April 2018

CPTR Newsletter – April 2018
In this issue:
- ReSeqTB to be Adopted for WHO Surveillance of Drug-Resistant TB
- New Collaborations Expand TB-PACTS Database and Leverage Data for Analyses
- CPTR Workgroup Works to Advance Regulatory Qualification of LAM, a Promising New Biomarker
- TB-PBPK Model Progresses Toward EMA and FDA Qualification
- TB-ReFLECT Update: A Collaborative Effort to Enhance TB Clinical Research
- New Project Launched to Evaluate Adaptive TB Clinical Trial Designs
- C-Path Partners with Catalysis Foundation for Health to Establish Database with Visualization and Analytic capabilities
—
ReSeqTB to be Adopted for WHO Surveillance of Drug-Resistant TB
The Relational Sequencing TB Data Platform (ReSeqTB), a global TB knowledge base for predicting TB drug resistance, will be adopted as the World Health Organization (WHO) bioinformatics platform for sequence-based surveillance of drug-resistant TB (DR-TB). In this new project phase, ReSeqTB will support surveillance and global policy development for sequencing-based clinical diagnosis of DR-TB. It could also serve as a model for the broader goals of establishing global antimicrobial resistance surveillance and clinical diagnosis networks for WHO high-priority pathogens. The development of ReSeqTB has been made possible through the collaboration of WHO, Foundation for Innovative New Diagnostics (FIND), and Critical Path Institute (C-Path) through their participation in the CPTR Initiative.
“Genetic sequencing is the future of surveillance and diagnosis of DR-TB,” said Dr. Tereza Kasaeva, Director of the WHO Global TB Programme. “With ReSeqTB, sequencing can be used to rapidly and accurately estimate prevalence of resistance to anti-TB drugs. This platform has the potential to expand our understanding of the genetic basis of drug resistance and open the way for the use of sequencing for effective clinical management of patients with DR-TB, saving millions of lives.”
“ReSeqTB is actively collecting, standardizing, and aggregating genomic, phenotypic and, when available, clinical data to support the development of global policy for use of sequencing-based diagnostics in high-burden settings” said Dr. Debra Hanna, Executive Director of C-Path’s Critical Path to TB Drug Regimens (CPTR) initiative. “Together with our partners, we are working with the US FDA to pursue regulatory clearance of ReSeqTB to support patient management decisions.”
ReSeqTB is changing the DR-TB landscape. “ReSeqTB is already informing the development of commercial TB molecular diagnostic solutions, and the confidence-graded mutations list is becoming a standard for interpretation of existing and future molecular diagnostics,” said Dr. Catharina Boehme, CEO of FIND. “Building genetic sequencing into global diagnostics policy is an exciting advance. Enhanced diagnostics mean that patients can get a diagnosis earlier, which means they can access the right treatment more quickly – and the transmission of DR-TB is curtailed.”
Read full press release here: https://cpathdev3.lotosnile.com/global-health-partners-accelerate-uptake-of-genetic-sequencing-for-surveillance-and-diagnosis-of-drug-resistant-tuberculosis/
—
New Collaborations Expand TB-PACTS Database and Leverage Data for Analyses
The TB PACTS Steering, including representatives from C-Path, the Special Programme for Research and Training in Tropical Diseases (TDR) at WHO, TB Alliance, St. George’s University of London, University of California, San Francisco, and the Innovative Medicines Initiative (IMI), were enthusiastically in favor of integrating datasets previously consolidated by the IMI PreDiCT-TB consortium into the TB-Platform for Aggregation of Clinical TB Studies (TB-PACTS). The consolidated IMI PreDiCT-TB consortium database currently contains 32 clinical data sets, most mapped to the CDISC TB 1.0 standard. To-date, TB-PACTS has received data transfers for thirteen of these datasets. An additional eight datasets are expected to be integrated into the platform within the year.
This collaboration will expand the aggregated Phase 3 TB clinical trial data in the TB-PACTS data platform that is publicly available to qualified researchers. Within CPTR, the data will be used to expand the ongoing modeling and simulation efforts (e.g., refine quantitative models linking dynamic changes in time to liquid culture positivity (TTP) to clinically relevant endpoints for Phase 3 trials such as culture conversion and durable cure).
“When we established TB-PACTS we did it not only to make data from TDR-sponsored studies available, but also (and mostly) to create a trusted system where others too would be confident to share the data they generated,” said Dr. Piero Olliaro, head of intervention and implementation research at TDR. “This is happening now with the IMI PreDiCT-TB datasets, and we look forward for others to follow, so that researchers can have as many data as possible to investigate and fill critical knowledge gaps in tuberculosis.”
—
CPTR Advances Regulatory Discussions on Promising Tools and Methodologies to Inform and Enhance Decision-making in TB Drug Development
CPTR Workgroup Works to Advance Regulatory Qualification of LAM, a Promising New Biomarker
CPTR’s Biomarker and Clinical Endpoint Workgroup continues to progress the investigation of lipoarabinomannan (LAM) as a non-culture based pharmacodynamic biomarker to assess treatment response in clinical trials evaluating novel TB drug candidates.
LAM is a major component of mycobacterium cell wall, comprising up to 1.5% of total bacterial weight. A new immunoassay (LAM-ELISA) has been developed by Otsuka Pharmaceutical Co, Ltd (Otsuka)., which uses monoclonal antibodies against novel, unique LAM epitopes with high sensitivity and specificity to quantify sputum LAM concentration. Under the proposed context of use, sputum LAM could serve as a potential real-time decision-making tool in adaptive clinical designs of new TB treatment regimens.
Building off a successful Critical Path Innovation Meeting with the FDA earlier in the year, CPTR received a positive response from FDA to the Letter of Intent submission, formally accepting the biomarker into FDA’s Qualification Program on October 31, 2017. The team has been advancing discussions with the EMA in parallel, with plans to submit documentation on the biomarker to both agencies in 2Q 2018.
TB-PBPK Model Progresses Toward EMA and FDA Qualification
CPTR and Simcyp have successfully developed a physiologically-based pharmacokinetic model for tuberculosis (TB-PBPK). This model incorporates relevant pathophysiological changes that occur in TB-infected individuals to describe drug distribution from the systemic circulation into the different lung compartments and tissues, including granulomas. It may be utilized for simulations with a variety of virtual populations, including a virtual South African population. This model is complete and fully deployed within the Simcyp® simulator environment. When applied to a library of standard of care (SOC) or emerging data for new compounds, this model may be used to inform early clinical development through predictions of drug distribution to the target tissues.
CPTR and Simcyp submitted a Letter of Intent and a briefing package to the EMA on November 22, 2017 to start the formal scientific advice qualification process for the PBPK model as a TB drug development tool. CPTR received formal feedback from EMA prior to a scheduled meeting with EMA’s Scientific Advice Working Party (SAWP) to discuss qualification on February 7, 2018. The EMA endorsed the updated context of use statement, to focus on the application of the PBPK platform to inform the design healthy volunteer studies for drugs that are not P-gp substrates, administered as monotherapy, for which plasma concentrations are known. Further refinements to the model are being investigated which will serve as the basis for a final qualification decision by EMA.
CPTR held an informal meeting with the FDA on December 1, 2017 to discuss next steps for regulatory interaction for the potential endorsement of the PBPK platform. Per FDA’s request, CPTR will initiate a Fit-For-Purpose submission to FDA, with a Letter of Intent and Briefing Book, incorporating the scientific advice from EMA.
Application of novel tools and methodologies can inform drug development decisions throughout the process:
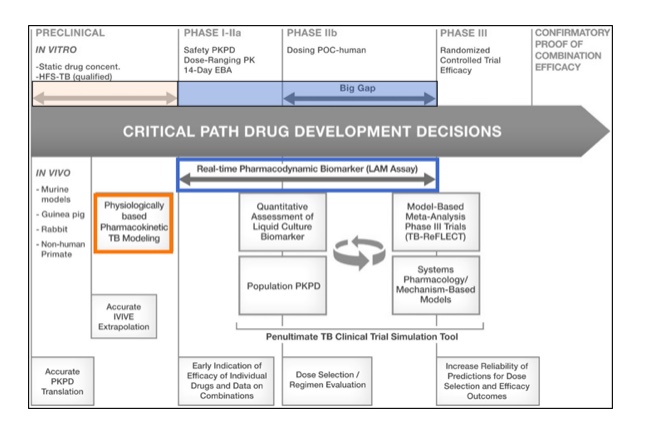
—
TB-ReFLECT Update: A Collaborative Effort to Enhance TB Clinical Research
In 2016, C-Path and the Global TB Programme of the World Health Organization (WHO) partnered with researchers from the University of California, San Francisco (UCSF), to develop leading-edge quantitative analyses of data from the TB-Platform for Aggregation of Clinical TB Studies (TB-PACTS) database. The objective of this collaboration, called TB reanalysis of fluoroquinolone clinical trials (TB-ReFLECT), is to extract from these analyses key lessons from the TB-PACTS platform, and then package such lessons as tools to optimize future TB trial design.
The TB-ReFLECT team completed sensitivity analyses in November 2017, which included different classifications of unfavorable outcomes, and efficacy analyses that examined random effects of pharmacokinetic (PK) imputations and parametric survival models. UCSF recently presented on TB-ReFLECT efficacy analyses at a WHO workshop on clinical design, and a manuscript detailing these analyses has been submitted for peer review.
The final stage of the project, safety analysis, is in progress and will analyze proportionality approaches to identify specific adverse events or clusters of adverse events, to inform benchmarking of the HRZE regimen in terms of safety.
CPTR will hold a face-to-face Steering Committee meeting to discuss how to message major learnings from TB-ReFLECT findings and discuss extensions to the project and/or integrations of the analyses into other CPTR projects.
—
New Project Launched to Evaluate Adaptive TB Clinical Trial Designs
CPTR’s Biomarker and Clinical Endpoint Workgroup (BCE-WG) launched a new effort to evaluate adaptive TB clinical trial designs, via simulation. This project aims to evaluate a set of Phase 2 and 3 clinical development strategies and compare their overall efficiency and performance in selecting potential pan-TB regimens. A Steering Committee, consisting of experts in the field from academia, NGOs, and industry, was created to develop the scope, metrics and quantitative work for this project. Quantitative analyses simulation work is scheduled to commence in 2018.
“Adaptive clinical trial designs using biomarkers to rapidly select the most promising treatments have been used with much success in other disease areas,” said Dr. Patrick Phillips, Professor, UCSF. “We need to be using the most efficient tools to rapidly develop safer and more efficacious TB regimens that can bring real patient benefit. In this project, we will develop and critically appraise some of the most promising biomarker-driven clinical development strategies, providing guidance to help developers speed novel regimens to market.”
—
C-Path Partners with Catalysis Foundation for Health to Establish Database with Visualization and Analytic capabilities
C-Path and Catalysis Foundation for Health (CFH) have partnered to share data from the clinical study “Mycobacterium tuberculosis (Mtb) Biomarkers for Diagnosis and Cure.” The collaboration seeks to identify biomarkers that correspond to detailed PET/CT information collected in this study, and to ultimately identify blood, urine, or saliva biomarker(s) that could replace sputum as a means of both diagnosing a patient and tracking their treatment progress.
Read more about the corresponding transcriptomics data for this study here:
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE89403.


